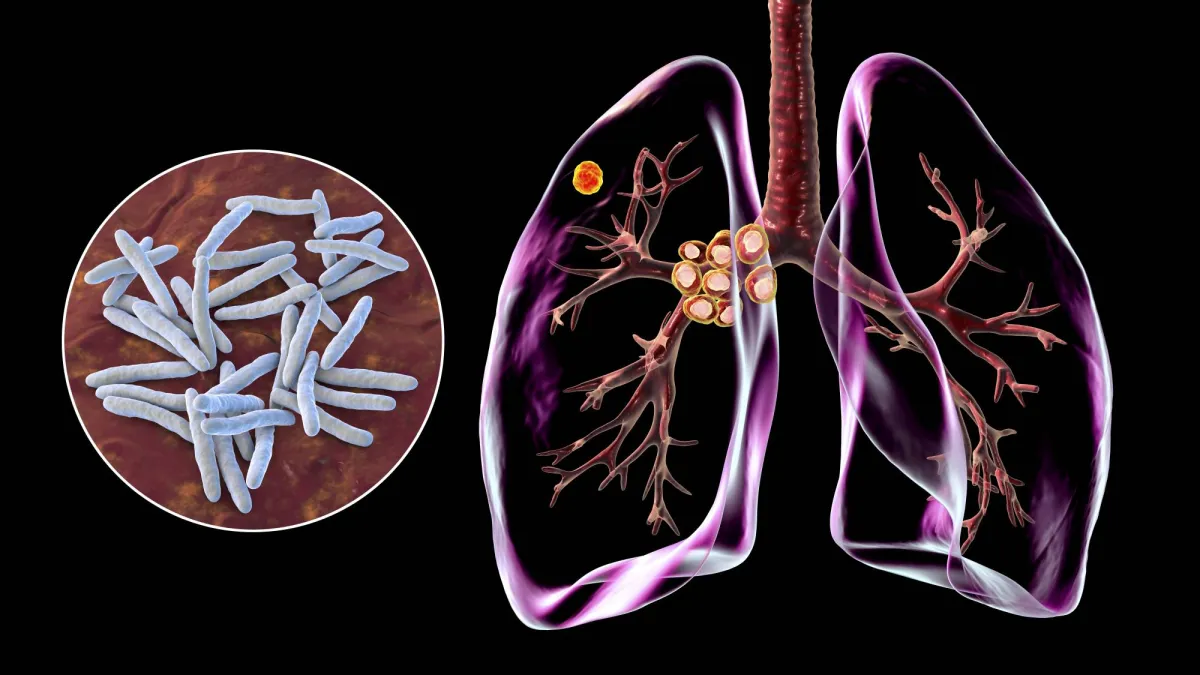
In a significant scientific advancement, a recent study from Spain has unveiled an innovative method for understanding the spread of tuberculosis through the use of genetic sequencing techniques to monitor infection transmission with unprecedented accuracy.
On the occasion of World Tuberculosis Day, this study emphasizes the increasing role of DNA analysis in uncovering hidden transmission chains of the disease and directing health interventions more effectively. In the Catalonia region, over 1200 tuberculosis cases are diagnosed annually, but the question remains: how many cases go undetected? How does the bacteria actually spread within communities?
Event Details
For the first time, researchers have widely employed genetic sequencing techniques to create an accurate map of tuberculosis spread across the region. The results, published in the journal "Frontiers in Microbiology" on March 20, 2026, reveal hotspots of the disease, as well as the genetic patterns of the circulating bacterial strains and the associated population groups.
This study is the result of a scientific collaboration between the Germans Trias i Pujol Research Institute and its university hospital, along with the Biomedical Research Institute in Valencia. It represents the first published results of the experimental TB-SEQ program. Launched in late 2021, this program aims to integrate genetic sequencing techniques into routine epidemiological surveillance systems for tuberculosis in Catalonia, marking a qualitative shift in the monitoring of infectious diseases.
Background & Context
Traditional methods of tracking tuberculosis rely on contact tracing, which involves asking infected patients about individuals they have spent time with and then testing those individuals. However, this method has blind spots, as individuals may not realize they have been exposed to the infection or may not remember every interaction.
Genetic sequencing offers a different lens; by analyzing the genetic material of the Mycobacterium tuberculosis bacteria from each patient, scientists can compare strains. If two patients have nearly identical bacterial genomes with only a few mutations separating them, it is highly likely they are part of the same transmission chain.
Impact & Consequences
The study also analyzed bacterial strains collected from across Catalonia between December 2021 and June 2023. The results show that the most common strain, known as "L4," is widespread throughout the region, found among both native populations and residents born outside Spain.
However, other strains tell a more specific story, as strains like L1/EAI, L2/Beijing, and L3/CAS frequently appear in certain geographic areas and are often linked to individuals originating from parts of the world where these subtypes are prevalent.
Regional Significance
Tuberculosis is often thought of as a disease of the past, but the numbers tell a different story. Globally, more than 10 million people contract tuberculosis each year, with approximately 1.5 million dying from the disease. In Catalonia, the incidence rate is around 15.2 new cases per 100,000 people, representing an ongoing public health challenge.
This current study provides a baseline—a genetic snapshot—of tuberculosis in Catalonia over the past 18 months. However, the true value of genomic surveillance lies in its continuous use. By sequencing new cases and comparing them to a growing database, public health officials can identify previously unknown transmission chains and halt outbreaks before they spread.